Abstract
Purpose of Review
Since the introduction of Archaea as a new domain of life more than 45 years ago, progress in their phylogenetic classification and knowledge of their exclusive biological characteristics has identified archaea as unique microorganisms which are widespread in extreme but also in various moderate ecosystems, including eukaryotic hosts. However, archaea are still neglected players within microbiomes, and research on archaea-bacteria interactions is still in its infancy due to methodological challenges.
Recent Findings
This review summarizes the current knowledge of archaea as components within microbiomes and focuses on their interactions with their bacterial neighbors and the principles of archaeal interactions.
Summary
Archaea are common constituents of animal and human microbiomes, which are dominated by Euryarchaeota. The gastrointestinal tract is the most studied body site, where archaea account for up to 4% of all microorganisms, primarily represented by methanogens. No archaeal pathogen has yet been identified, although methanogens are hypothesized to be indirectly involved in pathogenicity. Archaeal interactions comprise symbiotic relationships, and the cell membrane and wall might be as crucial as quorum sensing/quenching for these interactions. Particularly, syntrophic interactions under energy-deficiency stress seem to be an essential strategy for archaea. However, more research is urgently needed to discover how archaea sense their environment, compete with bacteria, and interact within complex microbiomes associated with multicellular organisms.
Similar content being viewed by others
Introduction
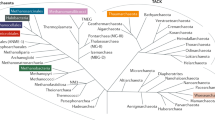
Archaea are prokaryotes belonging to the third domain of life, first classified separately from bacteria in 1977 by Carl Woese and George E. Fox based on their ribosomal RNA genes [1••, 2]. Although the classification of archaea is a rapidly progressing field, current classifications categorize this domain into four large clades, namely the Euryarchaeota, the TACK superphylum (Thaumarchaeota, Aigarchaeota, Crenarchaeota, Korarchaeota), the Asgard archaea, and the DPANN clade (Diapherotrites, Parvarchaeota, Aenigmarchaeota, Nanoarchaeota) [3, 4] (Fig. 1). Euryarchaeota represents the most extensively studied phylum as it includes the methanogens, which play an essential role in (syntrophic) anaerobic degradation processes [5] (Fig. 1). Besides this complex phylogeny, archaea harbor characteristics distinct from the other domains of life [6]. These unique characteristics are the lack of peptidoglycans in their cell wall and ether-linked lipids in their cell membrane. Archaea also show higher diversity in cell wall structures, e.g., surface layer proteins and heteroglycans [6]. Comparative analyses of archaeal genomes have identified several molecular conserved signatures uniquely present in archaea (l groups). One unique feature of archaea is the metabolic production of methane (methanogenesis) [6]. During the last decades, these exclusive properties were mainly regarded as unique adaptions to various extreme environments inhabited by archaea [7]. However, several studies have since discovered that archaea also live in moderate habitats, including soil and ocean [8]. They have been reported to be stable commensals of multicellular host organisms (plants and animals), where they participate in functions such as growth promotion, nutrient supply, protection against abiotic stress, methanogenesis, transformation of heavy metals, trimethylamine metabolism, and immune modulation (reviewed in [5]). Studies have confirmed the presence of archaea in various human body sites, such as the gut, skin, nose, mouth, lungs, and vagina [9,10,11]. Although enormous progress has already been made in archaeal research [12•], their unique physiological, structural, and molecular properties complicate further investigations, particularly in the context of microbiome research [13•, 14••]. This review aims to summarize the current knowledge about archaea as components within microbiomes and focuses on their interactions with their bacterial neighbors and the archaeal communication types used for these interactions.
Archaeal phylogeny. Schematic representation of a Maximum Likelihood phylogenetic tree based on 16S rRNA sequences from Archaea and Bacteria, adapted from [3]
Archaeal Interactions in Natural Ecosystems
Researchers only recently became aware of how archaea interact with each other and organisms of other domains. For instance, methanogens are involved in essential steps of the global methane cycle, partially conducted in a symbiotic interaction with herbivorous animals or sulfate-reducing bacteria [15]. Many syntrophic associations have been described for hydrogenotrophic methanogens, such as fermentative Acetobacterium and Syntrophobacter, Desulfovibrio, Thermoanaerobacter, Desulfotomaculum, and Pelotomaculum (reviewed in [15]). Besides microbial assemblages, archaea further interact with multicellular eukaryotes. Recent next-generation sequencing studies revealed highly abundant and diverse archaeal signatures in plant-associated microbiomes [16]. Particularly, plant roots and rhizospheres provide anoxic or oxygen-depleted micro-niches for methanogens and ammonium-oxidizing archaea [17]. Notably, plant genotypes affected the composition and structure of these archaeal communities [18]. The interaction could be based on syntrophic nitrogen cycling, and archaea are suggested to affect the nutrient exchange and coping with environmental stress [19]. Although there is initial evidence of specific interactions among archaea and between archaea and other domains of life, their mechanisms and ecological roles often remain unclear.
Archaea in Animals
Animals have been shown to be inhabited not only by trillions of bacteria and fungi but also by archaea, more precisely primarily Euryarchaeota, with a high abundance of anaerobically growing methanogens found within their digestive tracts [7]. Only a few studies have reported Crenarchaeota and Thaumarchaeota within animal microbiomes [20, 21]. The presence of archaea has been confirmed in sponges, mollusks, corals, arthropods, vertebrates, and humans (reviewed in [5, 15]). Investigations using Axinella sponge species associations with Cenarchaeum symbiosum or other filamentous Crenarchaeota [22], and the symbiosis of the demosponge Tentorium semisuberites with Cren- and Euryarchaeota [23], pointed to an archaeal contribution to the sponge nitrogen metabolism. A role within nitrogen cycling has also been identified in marine mollusks for Thaumarchaeota phylogenetically related to Nitrosopumilus maritimus [24]. Among arthropods, the largest animal phylum, exclusively methane-producing archaea were detected in millipedes, cockroaches, termites, and scarab beetles [25]. Here, the methanogens were primarily found in the hindguts of these animals [26]. In these interactions, archaea utilize hydrogen, carbon dioxide, and acetate resulting from the anaerobic degradation of lignocellulose [27].
Within the methanogenic archaea, particularly Methanobrevibacter species are extraordinarily well-adapted to interact with animal hosts and microbial neighbors [28••]. Consequently, Methanobrevibacter species are the predominant archaea in the gastrointestinal tracts (GITs) of various ruminants and non-ruminants, including cattle, sheep, reindeer, goats, buffalo, pigs, rhinoceroses, and chickens, among others (reviewed in [5, 28••]). In ruminants, archaea-bacteria consortia produce more than one-third of the methane [29]. Systematic analyses of methane production in the guts of over 250 vertebrates revealed that methane production and, thus, the presence of methanogens depends on the phylogenetic lineage [25]. Methanobrevibacter were shown to be flexible exponents of syntrophic interactions that enhance the efficiency of bacterial polysaccharide fermentation through methanogenesis since they can consume various fermentation products of primary and secondary fermenters, such as methanol, hydrogen, and carbon dioxide [30]. Methanobrevibacter smithii is known as a partner in a syntrophic interaction with Bacteroides thetaiotaomicron [31]. Here, the archaeon affected the expression of Bacteroides enzymes responsible for producing formate and acetate, which the archaeon utilizes. In gnotobiotic mice, the consortium was assumed to affect the energy balance of the host [32]. Furthermore, other methanogenic archaea, like Methanosphaera, Methanosarcina, Methanomassiliicoccus, and Methanimicrococcus, have been identified in the gut of various animals despite their low abundance [5]. Besides the archaeal colonization of the GIT, various animals also carry methanogens, Haloarchaea and Thaumarchaeota on the skin [33]. Also, those archaeal groups consume metabolic end products from the host and the microbiome and are supposed to affect the host [34••]. Although the archaeal community, also called archaeome, is now increasingly recognized as an essential component of (host-associated) microbiomes, contributing to methane production and potentially involved in disease-relevant processes, the underlying interaction mechanisms are still largely unknown [28••].
Role of Archaea in Humans
The human microbiome contains numerous archaea, particularly on the skin, in the respiratory tract, and in the GIT [10] (Fig. 2). Several studies estimated that approx. 500–1000 different microbial species are present within the human body at any time, accounting for 4 × 1013 cells. Archaea are predicted to account for around 1% of the total microbial cell count [35]. The human archeaome includes many lineages, including Methanobacteriales, Methanomassiliicoccales, Methanomicrobiales, Methanosarcinales, Halobacteriales, Thaumarchaeota (Nitrososphaeria), and members of the DPANN clade [28••](Fig. 1, 2). The oral cavity harbors various methanogenic strains of the genera Methanobrevibacter and Methanomassiliicoccus, often enriched in patients with periodontal disease [36]. Recent studies even identified halophilic and thermophilic Euryarchaeota and Thaumarchaeota in the human mouth [37] (Figs. 1, 2). The human skin is inhabited by Thaumarchaeota (Nitrososphaera), assumed to be responsible for ammonium turnover, and associated with odor reduction and skin improvement [33] (Fig. 2). Further, age and skin physiology affect the human skin archaeome [38]. Few studies describe the presence of archaea in human nasal, lung, and vaginal microbiomes [39••] (Fig. 2). As mentioned for animals, the vast majority of human-associated archaea is found in the intestine, accounting for up to 10% of the anaerobic community [40] (Fig. 2). The abundance of methanogenic archaea in the human gut is highly variable and represented by two physiological types of humans regarding gas exhalation (methane emitters and non-emitters) [41•]. While the specific role of non-methanogenic archaea in the human body remains to be explored, methanogens maintain numerous syntrophic relationships with resident bacteria by being involved in fermentation processes through hydrogen removal [28••]. M. smithii was the first archaeon isolated from human feces and is the most abundant archaeon found in the intestine [42]. M. smithii genomes displayed genomic adaptations to the human gut, such as producing surface glycans and adhesion-like proteins [30]. These adaptations are considered to originate from bacteria through lateral gene transfer [43]. Besides M. smithii, other methanogens such as Methanosphaera stadtmanae, and Methanomassiliicoccus luminyensis have been detected in human stool samples [44]. The use of modern high-throughput sequencing techniques revealed the presence of additional community members from the Eury- and Crenarchaeota, such as Desulfurococcales, Crenarchaeales, Sulfolobales, Thermoproteales, Archaeoglobales, Halobacteriales, Methanosarcinales, Methanobacteriales, Methanococcales, Methanopyrales, Thermococcales and Thermoplasmatales within the human intestine [45].
Although archaea can generally be considered commensal or beneficial organisms, and no archaeal pathogen is currently known, reports of archaeal involvement in human diseases are accumulating, particularly in immunocompromised individuals [46••]. Archaea have been found to have immunomodulatory abilities. For example, certain species of archaea can stimulate the immune system, increasing the production of cytokines and other immune cells [28••, 42, 47]. Other studies have suggested that archaea can exert anti-inflammatory effects by reducing the production of pro-inflammatory cytokines, such as methanogenic archaea, or increasing the production of anti-inflammatory cytokines [42, 48]. The exact mechanisms by which archaea exert their immunomodulatory effects are not yet fully understood, but it is thought that they may act through interactions with the gut microbiome and the gut epithelial cells [28••]. Other studies have proposed that archaea may interact with Toll-like receptors (TLRs) on immune cells, activating immune responses [49]. It is assumed that M. smithii may contribute to developing autoimmune diseases through various mechanisms, including producing metabolites, like methane, that activate immune cells and stimulate inflammation [34••]. In addition to its direct effects on the immune system, M. smithii may also indirectly contribute to autoimmune disease development by altering the gut microbiome and disrupting the balance of commensal microorganisms [50, 51]. Dysbiosis has been implicated in the development of various autoimmune diseases (reviewed in [28••]). Furthermore, M. stadtmanae was more abundant in the gut microbiome of patients with inflammatory bowel disease (IBD) [52] and may be associated with an increased risk of other autoimmune diseases, such as type 1 diabetes [53]. However, the effects of these interactions are not yet fully understood, and more research is needed to determine the precise ways archaea may affect the human immune system. It is also possible that different species of archaea may have different effects on the immune system and that the effects may depend on the context in which the interaction occurs.
Increasing evidence suggests that archaea may play a role in the development and progression of cancer in humans through various mechanisms, including the production of carcinogenic metabolites, modulation of immune responses, and alteration of the tumor microenvironment [54, 55]. Further, alterations in the abundance and diversity of archaeal populations may be associated with an increased risk of colorectal cancer [55]. For example, certain species of archaea, such as M. smithii, may be more abundant in the gut microbiome of individuals with colorectal cancer than in healthy individuals [56]. Similarly, hydrogen sulfide-producing archaea have been implicated in the development of gastric cancer due to the ability of hydrogen sulfide to cause DNA damage and inhibit apoptosis [57]. Other studies have suggested that archaea may be involved in oral infections and periodontitis [36, 58]. In particular, Methanobrevibacter oralis has been identified in the oral microbiomes of patients with periodontitis [59].
Principles of Archaea-Bacteria Interactions — Examples
Several types of symbiosis are known for microorganisms, including mutualism, commensalism, amensalism, parasitism, and predation [60]. These different types of symbiosis can have significant implications for microbial communities’ ecology and function and host organisms’ health and well-being. An example of a mutualistic relationship involving archaea, also called syntrophy, is the previously mentioned interaction between methanogenic archaea and anaerobic bacteria in the gut of ruminants [61] (Fig. 3, mutualism). Ruminants, such as cows, have a complex digestive system that allows them to break down rigid plant materials through fermentation. Methanogenic archaea are an essential part of this process, as they produce methane as a byproduct of the fermentation process. However, methane production is energetically costly, and the methanogens rely on other microbes in the gut to provide them with the necessary substrates for methane production. In exchange for these substrates, the methanogens provide the other microbes with an environment free of hydrogen gas, which is toxic [61, 62]. Moreover, the archaeon Acidianus and the bacterium Desulfurococcus coexist in acidic hot springs to perform sulfur cycling [63]. Acidianus oxidizes elemental sulfur to sulfuric acid, while Desulfurococcus reduces sulfate to sulfide. The interaction between these two microorganisms benefits both, as Acidianus provides Desulfurococcus with the sulfuric acid it needs to produce energy, and in turn, Desulfurococcus consumes the toxic sulfide produced by Acidianus. Within nitrogen cycling, the ammonia-oxidizing archaeon N. maritimus and the nitrite-oxidizing bacterium Nitrospira coexist in marine environments [64]. N. maritimus provides Nitrospira with the nitrite it needs to produce energy, and in turn, Nitrospira consumes the toxic nitrite produced by N. maritimus. Commensal symbiosis including archaea was shown in the guts of termites and humans, where one organism benefits from the association while the other is neither benefited nor harmed [65]. Similarly, methanogens at the surface of corals detoxify the environment [66](Fig. 3, commensalism), and in the soil, they degrade complex organic matter used by other microbes [67]. Amensalism describes a relationship with one partner negatively impacted by the presence of another organism while this organism is unaffected at all. One example of amensalism involving archaea can be again found in human guts [68]. M. smithii produces methane as a byproduct of its metabolism, while Desulfovibrio piger produces hydrogen sulfide as a byproduct. The presence of D. piger in the gut can reduce the sulfate levels available for M. smithii, thus inhibiting the growth and activity of the archaeon [30, 69] (Fig. 3, amensalism). Sulfur compounds are also essential for the survival of archaea and bacteria in deep-sea hydrothermal vents; consequently, the organisms compete with each other and get both negatively impacted [70]. For instance, Thermococcus can outcompete sulfur-reducing bacteria for sulfur compounds, using specialized enzymes that enable them to oxidize sulfur compounds to sulfate, releasing energy in the process [71] (Fig. 3, competition). Other archaea, such as Methanosarcina, can also outcompete sulfur-reducing bacteria by consuming hydrogen and carbon dioxide, which can limit the availability of these compounds for bacterial growth [72]. Archaea, like Methanosaeta harundinacea and Haloferax volcanii, compete with other microorganisms for space in their environment by forming biofilms and thus preventing the colonization of others [73]. There are currently no verified examples of archaea that are obligate parasites of other organisms. However, some archaea interact with other organisms in a way that can be considered to be parasitic [74]. For instance, some archaeal species have been found to live as endosymbionts in protists or amoebae [44]. M. luminyensis has been found in the ciliate protozoan Trimyema compressum [75], M. arboriphilicus in the amoeba Acanthamoeba polyphaga [76], and M. stadtmanae in the ciliate protozoan Metopus contortus [77]. A parasitic relationship exists between the thermoacidophilic archaeon Sulfolobus and the double-stranded DNA virus Sulfolobus Turreted Icosahedral Virus (STIV) [78] (Fig. 3, parasitism). STIV is an obligate parasite of Sulfolobus and depends on its host for replication and survival. STIV uses the host’s cellular machinery upon infection to replicate its genome, assemble new virus particles, and lyse the host cell to release the new virus particles [79]. The virus has evolved various mechanisms to manipulate the host’s cellular processes for its benefit [80]. For example, it encodes several proteins that can disrupt the host’s DNA repair mechanisms, thus preventing the host from repairing the DNA damage caused by the virus. STIV also encodes proteins that suppress the host’s immune response, allowing the virus to evade detection and destruction by the host’s defense mechanisms [80]. Archaea are generally not considered predators, lacking the specialized structures and mechanisms to capture and consume other microorganisms. However, predation of archaea is assumed to occur [81]. For example, Parcubacteria (also known as Candidate Division OD1) from soil ecosystems are known to feed on archaea, and some protists can engulf and digest archaea as a food source [82]. The dorvilleid polychaete, Ophryotrocha labronica, completes its life cycle by preying on two strains of Euryarchaeota — Haloferax and Halobacterium. Both archaea offered unique lipids to the polychaete [83](Fig. 3, predation). Lastly, archaea simply coexist within complex microbiomes with other microbes without direct consequences for the neighbors and the host [5] (Fig. 3, neutralism). These are just a few examples of the complex interactions between archaea and bacteria in microbiomes. The exact nature and importance of these interactions can vary depending on the specific microbiome and the types of microorganisms present.
Archaeal interactions. Interactions can be beneficial, neutral, or harmful for one or both partners and can take many forms, including mutualism (methanogenic archaea and anaerobic bacteria both benefit in the gut of ruminants), commensalism (methanogens at the surface of corals detoxify the environment), amensalism (Desulfovibrio piger reduces sulfate levels unavailable for Methanobrevibacter smithii), predation (Parcubacteria feed on archaea in soil ecosystems), parasitism (Sulfolobus Turreted Icosahedral Virus (STIV) is an obligate parasite of Sulfolobus), competition (archaea and bacteria compete for sulfur compounds in deep-sea hydrothermal vents), and neutralism (archaea coexist with other microbes). Created in BioRender.com
Communication of Archaea
Archaea can communicate with each other and other microorganisms in their environment through various mechanisms, including cell–cell contact, chemical signaling, and quorum sensing (QS) [84, 85] (Fig. 4). Archaea interact with their environment through their unique cell membranes that are composed of ether-branched chain lipids (isoprenoids), enabling to withstand extreme conditions such as high temperatures, high salinity, and low pH [86] (Fig. 4). Archaea also have unique cell walls that can vary in composition depending on the specific type of archaea. Structures like Pseudomurein, S-layers, sheaths, and Methanochondroitin provide structural support and protection and facilitate communication and exchange of nutrients between cells [87] (Fig. 4). Beyond, archaea have specialized structures to interact with their environment, communicate with other cells, and move in response to changing conditions [88] (Fig. 4), among those the archaellum, a whip-like structure similar to the bacterial flagellum, is used for movement [89]. Further, pili and fimbriae, hair-like structures, are involved in adhesion to surfaces and conjugation [88]. For instance, Methanococcus voltae has type IV pili that help it to attach to surfaces [90]; Methanocaldococcus jannaschii, produces long, filamentous pili, allowing them to interact with their environment (motility), form complex communities with other microorganisms (adhesion to surfaces and cells), and adapt to changing conditions (conjugation) [91]. H. volcanii, Sulfolobus spp., and Thermococcus spp. form pili for conjugation (also known as mating and fusion, respectively) to transfer genetic material between cells through direct contact [92]. Nanotubes, which are long, thin structures, connect Ignicoccus hospitalis cells and allow the exchange of cytoplasmic material for metabolic cooperation between cells [93]. Other archaea, such as Thermococcus and Pyrococcus, have thread-like structures for attaching to surfaces or for movement [94], while some thermophilic and hyperthermophilic archaea, including members of the Pyrodictium and Ignicoccus, form cannulae. Cannulae are hollow, needle-like structures that connect neighboring cells for exchanging materials and information between them [95]. As unique long, flexible, grappling hook-like structure, Nanoarchaeum equitans has hami, used to attach to the surface of Ignicoccus [96]. Archaeal species, such as H. volcanii and Sulfolobus acidocaldarius can further produce small membrane vesicles, spherical structures that bud off the cell membrane [97]. These vesicles can contain a variety of molecules, including enzymes and signaling molecules facilitating nutrient uptake, communication, and defense against environmental stressors.
Chemical signals, such as (small) molecules or gases, are released by archaea into the environment and detected by other microorganisms in the vicinity, allowing for coordinated responses to changing environmental conditions [84]. Some of the most common chemical signals archaea use include metabolites that can influence the growth and behavior of other microorganisms [84]. For example, some methanogenic archaea produce acetate, which other microbes in the syntrophic relationship can use [98]. Similarly, extracellular enzymes are exchanged in syntrophic interactions [99]. Archaea can also use electron shuttles to interact with other microbes. Bacteria provide methanogenic archaea with electron donors, and the archaea provide the bacteria with a stable environment and a sink for the end-products of their metabolism [100]. Extracellular polymeric substances (EPS) produced by biofilm-forming archaea are other chemical signals in the attachment of cells to surfaces, each other, and other microbes [101]. EPS can play various roles, including cell attachment, nutrient uptake, and protection against environmental stress. Some archaeal species, such as the halophilic archaeon Halobacterium salinarum, produce EPS to survive in high-salt environments [102]. Another way archaea communicate is through quorum sensing (QS), a process by which microorganisms can detect the density of other microorganisms in their environment and adjust their behavior accordingly [85] (see Excursion – Quorum sensing in Archaea). Overall, the mechanisms by which archaea communicate in microbiomes are still not well understood, and further research is needed to elucidate the specific signaling pathways and molecules involved. However, communication and cooperation among various microorganisms are critical for the formation and function of microbial communities in their environment [8].
Excursion — Quorum Sensing in Archaea
Archaea use QS for cell-to-cell communication [85]. QS is a process by which microorganisms detect the density of their population and regulate gene expression as a community [103]. This can be beneficial for microbial survival and growth, as well as for interactions with host organisms and other microbiome members [104]. It typically involves producing, releasing, and detecting small signaling molecules called autoinducers. When the concentration of autoinducers in the environment reaches a certain threshold, it triggers a response that can result in changes in gene expression, behavior, or metabolism [103]. Overall, the use of QS among archaea is less well studied than in bacteria, but recent research has shed more light on the phenomenon in these organisms [85]. Several types of signaling molecules have been identified in archaea, including acyl-homoserine lactones (AHLs) [85]. These are signaling molecules commonly used by Gram-negative bacteria [105•]. However, AHLs have also been found in some archaea, such as Methanococcus maripaludis [106]. Recently, 11 new archaeal isolates with AHL-like activity against the LuxR-based AHL biosensor, including thermophiles, were identified [85, 107, 108]. Multiple AHL-like signal molecules were detected in Haloferax mucosum, Halorubrum kocurii, Natronococcus occultus, and H. salinarium [85]. A modification of AHL signaling molecules by a carboxyl group seems unique to archaea [106], and in M. harudinacea carboxyl-AHLs are assumed for one-way inter-domain cross-talk [106]. The work on M. harundinacea also described a putative AHL synthase, FilI, a histidine kinase different from any known bacterial AHL synthase [106]. Diketopiperazines (DKPs), known as signaling molecules in the bacteria Pseudomonas aeruginosa, Bacillus subtilis, and Streptomyces sp., have also been detected in archaea [109]. DKPs were suggested for regulating protease activity in Natronoccocus occultus [109]. Autoinducer-2 (AI-2) was shown to be involved in inter-species communication and biofilm formation in some archaea, e.g., Methanosarcina acetivorans, M. maripaludis, and Sulfolobus solfataricus [110•]. Recent research suggests that QS is crucial in many ecological and physiological processes. The direct evidence for QS in archaea is rare, but many phenotypes commonly regulated by QS in bacteria, such as biofilm formation, extracellular enzyme production, and membrane vesicle formation, are also present in archaea, suggesting a similar QS regulation of those phenotypes in archaea (reviewed in [85]). For instance, a biofilm phenotype associated with AHL activity was described for H. lacusprofundi [107]. M. jannaschii produces a phenotypic response when cultured with the bacterium Thermys maritima, potentially relying on QS molecules [111]. Co-cultured T. maritima and Pyrococcus furiosus produce AI-2 [111]. Moreover, a potential QS signal was observed in the supernatant from a Natrialba magadii culture involved in extracellular enzyme production [112].
Some evidence suggests that quorum quenching (QQ) may be present in certain archaeal species [85]. QQ is the process of interfering with QS in microorganisms, often through enzymes or small molecules [113]. In the environment, these QQ systems enable the turnover of QS signals in communities allowing for effective change between phenotypes or offering competitive advantages to some organisms by interference with these systems [114]. While QQ has been extensively studied in bacteria, it is less understood in archaea [85]. In S. solfataricus and S. acidocaldarius, the phosphotriesterase-like lactonases SsoPox and SacPox were identified, respectively, which inhibit short-chain (C4-HSL) and long-chain (3-oxo-C12-HSL) AHLs produced by the opportunistic pathogen P. aeruginosa PAO1 [115]. Further, DKPs synthesized by the haloarchaeon Haloterrigena hispanica might act as AHL mimics, blocking AHL detection of neighboring bacteria [109]. Further research is needed to fully understand the prevalence and significance of QS and QQ in archaea.
Conclusion
Recent research has highlighted the importance of communication between archaea and bacteria during microbe-microbe interactions, particularly in the context of symbiotic relationships within animal and human microbiomes. While bacteria have long been recognized as key players in microbiome ecology and function, emerging evidence suggests that archaea also play essential roles in these complex microbial communities. Studies have shown that archaea and bacteria can exchange chemical signals, such as QS molecules, to coordinate their activities and respond to environmental changes. In particular, symbiotic archaea have been found to play important roles in host digestion, metabolism, and immune function, highlighting the importance of understanding archaea-bacteria communication for human health. Overall, the growing recognition of archaea as key players in microbial communities underscores the need for further research into the mechanisms and consequences of archaea-bacteria communication during microbe-microbe interactions.
Data Availability
The data used in this review article are derived from previously published studies and publicly available sources. No new primary data were collected for this review. All cited references are provided within the manuscript, and readers are encouraged to consult these sources for further information.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
•• Baker BJ, De Anda V, Seitz KW, Dombrowski N, Santoro AE, Lloyd KG. Diversity, ecology and evolution of Archaea. Nature Microbiol. 2020;5(7):887–900. The paper provides an overview of the diversity, ecology, and evolution of archaea, highlighting recent advances in genomic and metagenomic techniques. The authors discuss the key features that distinguish archaea from bacteria and eukaryotes and explores the various ecological niches that archaea occupy. The authors highlight the important roles that archaea play in global nutrient cycling and discusses the evolutionary history of archaea.
Woese CR, Fox GE. Phylogenetic structure of the prokaryotic domain: the primary kingdoms. Proc Natl Acad Sci. 1977;74(11):5088–90.
Medina-Chávez NO, Travisano M. Archaeal Communities: the microbial phylogenomic frontier. Front Genet. 2022;12:693193.
Rinke C, Chuvochina M, Mussig AJ, Chaumeil P-A, Davín AA, Waite DW, et al. A standardized archaeal taxonomy for the Genome Taxonomy Database. Nat Microbiol. 2021;6(7):946–59.
Moissl-Eichinger C, Pausan M, Taffner J, Berg G, Bang C, Schmitz RA. Archaea are interactive components of complex microbiomes. Trends Microbiol. 2018;26(1):70–85.
Cavicchioli R. Archaea: molecular and cellular biology. In: Cavicchioli R, editor. Washington, USA: ASM Press; 2007. p. 523.
Bang C, Schmitz RA. Archaea: forgotten players in the microbiome. Emerg Topics Life Sci. 2018;2(4):459–68.
Flemming H-C, Wuertz S. Bacteria and archaea on Earth and their abundance in biofilms. Nat Rev Microbiol. 2019;17(4):247–60.
Belmok A, de Cena J, Kyaw C, Damé-Teixeira N. The oral archaeome: a scoping review. J Dent Res. 2020;99(6):630–43.
Koskinen K, Pausan MR, Perras AK, Beck M, Bang C, Mora M, et al. First insights into the diverse human archaeome: specific detection of archaea in the gastrointestinal tract, lung, and nose and on skin. MBio. 2017;8(6):e00824-e917.
Bang C, Schmitz RA. Archaea associated with human surfaces: not to be underestimated. FEMS Microbiol Rev. 2015;39(5):631–48.
• Sun Y, Liu Y, Pan J, Wang F, Li M. Perspectives on cultivation strategies of archaea. Microbial Ecol. 2020;79:770–84. The paper reviews various cultivation strategies that have been developed to overcome the challenges of cultivating archaea, including approaches such as co-cultivation with other microorganisms, the use of specialized growth media and culture conditions, and high-throughput screening methods. The authors also discuss the potential applications of cultivated archaea in biotechnology, including the production of biofuels, biopolymers, and other useful compounds.
• Forterre P. Archaea: a goldmine for molecular biologists and evolutionists. Archaea: Methods and Protocols. Springer; 2022; 1–21. The chapter published in the book "Archaea: Methods and Protocols" in 2022 by Springer, provides an overview of the unique features and potential applications of archaea in molecular biology and evolutionary studies. The author highlights the importance of comparative genomics for understanding the evolutionary relationships between different groups of organisms and for identifying new molecular targets for biotechnological applications. The chapter also discusses the potential applications of archaea in biotechnology.
•• Hoegenauer C, Hammer HF, Mahnert A, Moissl-Eichinger C. Methanogenic archaea in the human gastrointestinal tract. Nature Reviews Gastroenterology & Hepatology. 2022:1–9. The paper reviews recent studies on the diversity and abundance of methanogenic archaea in the human gut microbiota, as well as their potential interactions with other gut microbes and the host. The authors also discuss the possible links between methanogenic archaea and various gastrointestinal disorders, including constipation, inflammatory bowel disease, and colon cancer. The paper highlights the need for further research to fully understand the role of methanogenic archaea in the gut microbiota and their potential implications for human health.
Wrede C, Dreier A, Kokoschka S, Hoppert M. Archaea in symbioses. Archaea. 2012;2012:596846.
Buee M, De Boer W, Martin F, Van Overbeek L, Jurkevitch E. The rhizosphere zoo: an overview of plant-associated communities of microorganisms, including phages, bacteria, archaea, and fungi, and of some of their structuring factors. Springer; 2009.
He Y, Hu W, Ma D, Lan H, Yang Y, Gao Y. Abundance and diversity of ammonia-oxidizing archaea and bacteria in the rhizosphere soil of three plants in the Ebinur Lake wetland. Can J Microbiol. 2017;63(7):573–82.
Lee S-H, Kim S-Y, Ding W, Kang H. Impact of elevated CO 2 and N addition on bacteria, fungi, and archaea in a marsh ecosystem with various types of plants. Appl Microbiol Biotechnol. 2015;99:5295–305.
Valentine DL. Adaptations to energy stress dictate the ecology and evolution of the Archaea. Nat Rev Microbiol. 2007;5(4):316–23.
Turque AS, Batista D, Silveira CB, Cardoso AM, Vieira RP, Moraes FC, et al. Environmental shaping of sponge associated archaeal communities. PLoS One. 2010;5(12):e15774.
Durand L, Zbinden M, Cueff-Gauchard V, Duperron S, Roussel EG, Shillito B, et al. Microbial diversity associated with the hydrothermal shrimp Rimicaris exoculata gut and occurrence of a resident microbial community. FEMS Microbiol Ecol. 2009;71(2):291–303.
Margot H, Acebal C, Toril E, Amils R, Fernandez PJ. Consistent association of crenarchaeal Archaea with sponges of the genus Axinella. Mar Biol. 2002;140:739–45.
Pape T, Hoffmann F, Queric N-V, von Juterzenka K, Reitner J, Michaelis W. Dense populations of Archaea associated with the demosponge Tentorium semisuberites Schmidt, 1870 from Arctic deep-waters. Polar Biol. 2006;29:662–7.
Vortsepneva E, Chevaldonné P, Klyukina A, Naduvaeva E, Todt C, Zhadan A, et al. Microbial associations of shallow-water Mediterranean marine cave Solenogastres (Mollusca). PeerJ. 2021;9:e12655. https://doi.org/10.7717/peerj.12655.
Hackstein JHP, van Alen TA. Methanogens in the gastro-intestinal tract of animals. In: Hackstein JHP, editor. (Endo)symbiotic Methanogenic Archaea. Berlin, Heidelberg: Springer Berlin Heidelberg; 2010; 115–42.
Hackstein JH, Stumm CK. Methane production in terrestrial arthropods. Proc Natl Acad Sci U S A. 1994;91(12):5441–5. https://doi.org/10.1073/pnas.91.12.5441.
Brauman A, Doré J, Eggleton P, Bignell D, Breznak JA, Kane MD. Molecular phylogenetic profiling of prokaryotic communities in guts of termites with different feeding habits. FEMS Microbiol Ecol. 2001;35(1):27–36. https://doi.org/10.1111/j.1574-6941.2001.tb00785.x.
•• Borrel G, Brugère J-F, Gribaldo S, Schmitz RA, Moissl-Eichinger C. The host-associated archaeome. Nature Rev Microbiol. 2020;18(11):622–36. https://doi.org/10.1038/s41579-020-0407-y. The authors review recent advances in our understanding of the archaeal microbiome, focusing on the various host-associated environments where archaea have been identified, including the gut, oral cavity, skin, and genital tract. They discuss the diversity of archaeal taxa that have been identified in these environments, and the potential roles that they may play in host physiology and disease.
Finlay BJ, Esteban G, Clarke KJ, Williams AG, Embley TM, Hirt RP. Some rumen ciliates have endosymbiotic methanogens. FEMS Microbiol Lett. 1994;117(2):157–61. https://doi.org/10.1111/j.1574-6968.1994.tb06758.x.
Samuel BS, Hansen EE, Manchester JK, Coutinho PM, Henrissat B, Fulton R, et al. Genomic and metabolic adaptations of Methanobrevibacter smithii to the human gut. Proc Natl Acad Sci. 2007;104(25):10643–8.
Nkamga VD, Lotte R, Roger PM, Drancourt M, Ruimy R. Methanobrevibacter smithii and Bacteroides thetaiotaomicron cultivated from a chronic paravertebral muscle abscess. Clin Microbiol Infect. 2016;22(12):1008–9. https://doi.org/10.1016/j.cmi.2016.09.007.
Samuel BS, Shaito A, Motoike T, Rey FE, Backhed F, Manchester JK, et al. Effects of the gut microbiota on host adiposity are modulated by the short-chain fatty-acid binding G protein-coupled receptor, Gpr41. Proc Natl Acad Sci. 2008;105(43):16767–72. https://doi.org/10.1073/pnas.0808567105.
Probst AJ, Auerbach AK, Moissl-Eichinger C. Archaea on human skin. PLoS One. 2013;8(6):e65388. https://doi.org/10.1371/journal.pone.0065388.
•• Mafra D, Ribeiro M, Fonseca L, Regis B, Cardozo LF, Dos Santos HF, et al. Archaea from the gut microbiota of humans: Could be linked to chronic diseases? Anaerobe. 2022:102629. The paper reviews recent studies on the diversity and abundance of archaea in the human gut microbiota, as well as their potential interactions with other gut microbes and the host. The authors also discuss the possible links between archaea and chronic diseases such as inflammatory bowel disease, colorectal cancer, and metabolic disorders. The paper highlights the need for further research to fully understand the role of archaea in the gut microbiota and their potential implications for human health.
Zhu B, Wang X, Li L. Human gut microbiome: the second genome of human body. Protein Cell. 2010;1(8):718–25. https://doi.org/10.1007/s13238-010-0093-z.
Lepp PW, Brinig MM, Ouverney CC, Palm K, Armitage GC, Relman DA. Methanogenic Archaea and human periodontal disease. Proc Natl Acad Sci. 2004;101(16):6176–81. https://doi.org/10.1073/pnas.0308766101.
Dame-Teixeira N, de Cena JA, Côrtes DA, Belmok A, dos Anjos Borges LG, Marconatto L, et al. Presence of Archaea in dental caries biofilms. Arch Oral Biol. 2020;110:104606. https://doi.org/10.1016/j.archoralbio.2019.104606.
Moissl-Eichinger C, Probst AJ, Birarda G, Auerbach A, Koskinen K, Wolf P, et al. Human age and skin physiology shape diversity and abundance of Archaea on skin. Sci Rep. 2017;7(1):4039. https://doi.org/10.1038/s41598-017-04197-4.
•• Geesink P, Ettema TJG. The human archaeome in focus. Nature Microbiol. 2022;7(1):10–1. https://doi.org/10.1038/s41564-021-01031-6. The authors discuss recent studies that have used metagenomic sequencing to identify archaeal taxa in various human body sites. They highlight the diverse and often poorly characterized archaeal taxa that have been identified in these studies, and the potential roles that they may play in human health and disease. The paper also discusses the challenges associated with studying the human archaeome.
Eckburg PB, Bik EM, Bernstein CN, Purdom E, Dethlefsen L, Sargent M, et al. Diversity of the human intestinal microbial flora. Science. 2005;308(5728):1635–8. https://doi.org/10.1126/science.1110591.
•Chibani CM, Mahnert A, Borrel G, Almeida A, Werner A, Brugère J-F, et al. A catalogue of 1,167 genomes from the human gut archaeome. Nature Microbiology. 2022;7(1):48–61. doi: https://doi.org/10.1038/s41564-021-01020-9.The authors used metagenomic sequencing to analyze the archaeal diversity in fecal samples from over 1,000 individuals from various geographic locations and populations. They identified over 1,100 archaeal genomes, including many previously unknown species, and performed a detailed phylogenetic analysis to classify these genomes into distinct clades. The paper also provides insights into the functional potential of the human gut archaeome, identifying genes and pathways related to various metabolic processes and highlighting the potential roles of archaea in nutrient cycling and other gut functions.
Bang C, Weidenbach K, Gutsmann T, Heine H, Schmitz RA. The intestinal archaea Methanosphaera stadtmanae and Methanobrevibacter smithii activate human dendritic cells. PLoS One. 2014;9(6):e99411. https://doi.org/10.1371/journal.pone.0099411.
Lurie-Weinberger MN, Peeri M, Gophna U. Contribution of lateral gene transfer to the gene repertoire of a gut-adapted methanogen. Genomics. 2012;99(1):52–8. https://doi.org/10.1016/j.ygeno.2011.10.005.
Dridi B, Raoult D, Drancourt M. Archaea as emerging organisms in complex human microbiomes. Anaerobe. 2011;17(2):56–63.
Nam Y-D, Chang H-W, Kim K-H, Roh SW, Kim M-S, Jung M-J, et al. Bacterial, archaeal, and eukaryal diversity in the intestines of Korean people. J Microbiol. 2008;46:491–501.
•• Mohammadzadeh R, Mahnert A, Duller S, Moissl-Eichinger C. Archaeal key-residents within the human microbiome: characteristics, interactions and involvement in health and disease. Current Opinion in Microbiology. 2022;67:102146. The authors discuss the characteristics of archaea that make them well-suited for life in extreme environments, as well as their metabolic diversity and ability to interact with other microorganisms. They also highlight recent studies that have identified specific archaeal taxa that are consistently present in the human microbiome across different populations and geographic regions. The paper also discusses the potential roles of archaea in human health and disease, including their involvement in gut homeostasis and nutrient cycling, as well as their potential to contribute to disease states such as inflammatory bowel disease and colorectal cancer.
Cavicchioli R, Curmi PM, Saunders N, Thomas T. Pathogenic archaea: do they exist? BioEssays. 2003;25(11):1119–28.
Bang C, Vierbuchen T, Gutsmann T, Heine H, Schmitz RA. Immunogenic properties of the human gut-associated archaeon Methanomassiliicoccus luminyensis and its susceptibility to antimicrobial peptides. PLoS One. 2017;12(10):e0185919.
Vierbuchen T, Bang C, Rosigkeit H, Schmitz RA, Heine H. The human-associated archaeon Methanosphaera stadtmanae is recognized through its RNA and induces TLR8-dependent NLRP3 inflammasome activation. Front Immunol. 2017;8:1535.
Grine G, Drouet H, Fenollar F, Bretelle F, Raoult D, Drancourt M. Detection of Methanobrevibacter smithii in vaginal samples collected from women diagnosed with bacterial vaginosis. Eur J Clin Microbiol Infect Dis. 2019;38:1643–9.
Sogodogo E, Fellag M, Loukil A, Nkamga VD, Michel J, Dessi P, et al. Nine cases of methanogenic archaea in refractory sinusitis, an emerging clinical entity. Front Public Health. 2019;7:38.
Chen L, Wang J. Gut microbiota and inflammatory bowel disease. WIREs Mech Dis. 2022;14(2):e1540.
Traversi D, Scaioli G, Rabbone I, Carletto G, Ferro A, Franchitti E, et al. Gut microbiota, behavior and nutrition after type 1 diabetes diagnosis: a longitudinal study for supporting data in the metabolic control. Front Nutr. 2022;1–11.
Cai M, Kandalai S, Tang X, Zheng Q. Contributions of human-associated archaeal metabolites to tumor microenvironment and carcinogenesis. Microbiol Spectrum. 2022;10(2):e02367-e2421.
Coker OO, Wu WKK, Wong SH, Sung JJ, Yu J. Altered gut archaea composition and interaction with bacteria are associated with colorectal cancer. Gastroenterology. 2020;159(4):1459-70.e5.
Mira-Pascual L, Cabrera-Rubio R, Ocon S, Costales P, Parra A, Suarez A, et al. Microbial mucosal colonic shifts associated with the development of colorectal cancer reveal the presence of different bacterial and archaeal biomarkers. J Gastroenterol. 2015;50:167–79.
Ishaq SL, Moses PL, Wright A-DG. The pathology of methanogenic archaea in human gastrointestinal tract disease. Gut Microbiome Implic Hum Dis. 2016.
Matarazzo F, Ribeiro AC, Feres M, Faveri M, Mayer MPA. Diversity and quantitative analysis of Archaea in aggressive periodontitis and periodontally healthy subjects. J Clin Periodontol. 2011;38(7):621–7.
Bringuier A, Khelaifia S, Richet H, Aboudharam G, Drancourt M. Real-time PCR quantification of Methanobrevibacter oralis in periodontitis. J Clin Microbiol. 2013;51(3):993–4.
Martin BD, Schwab E. Symbiosis:“Living together” in chaos. Stud History Biol. 2012;4(4):7–25.
Morgavi D, Forano E, Martin C, Newbold CJ. Microbial ecosystem and methanogenesis in ruminants. Animal. 2010;4(7):1024–36.
Leahy SC, Janssen PH, Attwood GT, Mackie RI, McAllister TA, Kelly WJ. Electron flow: key to mitigating ruminant methanogenesis. Trends Microbiol. 2022;30(3):209–12.
Counts JA, Willard DJ, Kelly RM. Life in hot acid: a genome-based reassessment of the archaeal order sulfolobales. Environ Microbiol. 2021;23(7):3568–84.
Foesel BU, Gieseke A, Schwermer C, Stief P, Koch L, Cytryn E, et al. Nitrosomonas Nm143-like ammonia oxidizers and Nitrospira marina-like nitrite oxidizers dominate the nitrifier community in a marine aquaculture biofilm. FEMS Microbiol Ecol. 2008;63(2):192–204.
Bäckhed F, Ley RE, Sonnenburg JL, Peterson DA, Gordon JI. Host-bacterial mutualism in the human intestine. Science. 2005;1915–20.
Siboni N, Ben-Dov E, Sivan A, Kushmaro A. Geographic specific coral-associated ammonia-oxidizing archaea in the northern Gulf of Eilat (Red Sea). Microb Ecol. 2012;64:18–24.
Krzmarzick MJ, Taylor DK, Fu X, McCutchan AL. Diversity and niche of archaea in bioremediation. Archaea. 2018;2018.
Traore SI, Khelaifia S, Armstrong N, Lagier J, Raoult D. Isolation and culture of Methanobrevibacter smithii by co-culture with hydrogen-producing bacteria on agar plates. Clin Microbiol Infect. 2019;25(12):1561.e1-1561.e5.
Rey FE, Faith JJ, Bain J, Muehlbauer MJ, Stevens RD, Newgard CB, et al. Dissecting the in vivo metabolic potential of two human gut acetogens. J Biol Chem. 2010;285(29):22082–90.
Sievert SM, Kiene RP, Schulz-Vogt HN. The sulfur cycle. Oceanography. 2007;20(2):117–23.
Adams MM, Hoarfrost AL, Bose A, Joye SB, Girguis PR. Anaerobic oxidation of short-chain alkanes in hydrothermal sediments: potential influences on sulfur cycling and microbial diversity. Front Microbiol. 2013;4:110.
Zhang D, Vahala R, Wang Y, Smets BF. Microbes in biological processes for municipal landfill leachate treatment: community, function and interaction. Int Biodeterior Biodegradation. 2016;113:88–96.
Sekiguchi Y, Kamagata Y, Nakamura K, Ohashi A, Harada H. Fluorescence in situ hybridization using 16S rRNA-targeted oligonucleotides reveals localization of methanogens and selected uncultured bacteria in mesophilic and thermophilic sludge granules. Appl Environ Microbiol. 1999;65(3):1280–8.
Moissl-Eichinger C, Huber H. Archaeal symbionts and parasites. Curr Opin Microbiol. 2011;14(3):364–70.
Shinzato N, Watanabe I, Meng X-Y, Sekiguchi Y, Tamaki H, Matsui T, et al. Phylogenetic analysis and fluorescence in situ hybridization detection of archaeal and bacterial endosymbionts in the anaerobic ciliate Trimyema compressum. Microb Ecol. 2007;54:627–36.
Tokura M, Ohkuma M, Kudo T. Molecular phylogeny of methanogens associated with flagellated protists in the gut and with the gut epithelium of termites. FEMS Microbiol Ecol. 2000;33(3):233–40.
van Hoek AH, van Alen TA, Sprakel VS, Leunissen JA, Brigge T, Vogels GD, et al. Multiple acquisition of methanogenic archaeal symbionts by anaerobic ciliates. Mol Biol Evol. 2000;17(2):251–8.
Khayat R, Fu C-y, Ortmann AC, Young MJ, Johnson JE. The architecture and chemical stability of the archaeal Sulfolobus turreted icosahedral virus. J Virol. 2010;84(18):9575–83.
Brumfield SK, Ortmann AC, Ruigrok V, Suci P, Douglas T, Young MJ. Particle assembly and ultrastructural features associated with replication of the lytic archaeal virus Sulfolobus turreted icosahedral virus. J Virol. 2009;83(12):5964–70.
Ortmann AC, Brumfield SK, Walther J, McInnerney K, Brouns SJ, Van De Werken HJ, et al. Transcriptome analysis of infection of the archaeon Sulfolobus solfataricus with Sulfolobus turreted icosahedral virus. J Virol. 2008;82(10):4874–83.
Husnik F, Tashyreva D, Boscaro V, George EE, Lukeš J, Keeling PJ. Bacterial and archaeal symbioses with protists. Curr Biol. 2021;31(13):R862–77.
Seyler LM, Tuorto S, McGuinness LR, Gong D, Kerkhof LJ. Bacterial and archaeal specific-predation in the North Atlantic Basin. Front Mar Sci. 2019;6:555.
Thurber AR, Levin LA, Orphan VJ, Marlow JJ. Archaea in metazoan diets: implications for food webs and biogeochemical cycling. ISME J. 2012;6(8):1602–12.
Witzany G. Introduction: keylevels of biocommunication of archaea, in biocommunication of archaea. In: Witzany G, editor. Springer International Publishing: Cham; 2017. p. 1–16.
Charlesworth JC, Beloe C, Watters C, Burns BP. Quorum sensing in archaea: recent advances and emerging directions. Biocommun Archaea. 2017;119–32.
Caforio A, Driessen AJ. 2017 Archaeal phospholipids: structural properties and biosynthesis. Biochimica et Biophysica Acta (BBA)-Mol Cell Biol Lipids. 1862;11:1325–39.
Klingl A, Pickl C, Flechsler J. Archaeal cell walls. Bacterial Cell Walls Membr. 2019;471–93.
Ng SY, Zolghadr B, Driessen AJ, Albers S-V, Jarrell KF. Cell surface structures of archaea. J Bacteriol. 2008;190(18):6039–47.
Albers S-V, Jarrell KF. The archaellum: an update on the unique archaeal motility structure. Trends Microbiol. 2018;26(4):351–62.
Fonseca DR, Halim MFA, Holten MP, Costa KC. Type IV-like pili facilitate transformation in naturally competent archaea. J Bacteriol. 2020;202(21):e00355-e420.
Jarrell KF, Ding Y, Nair DB, Siu S. Surface appendages of archaea: structure, function, genetics and assembly. Life. 2013;3(1):86–117.
Wagner A, Whitaker RJ, Krause DJ, Heilers J-H, Van Wolferen M, Van Der Does C, et al. Mechanisms of gene flow in archaea. Nat Rev Microbiol. 2017;15(8):492–501.
Marguet E, Gaudin M, Gauliard E, Fourquaux I, le Blond du Plouy S, Matsui I, et al. Membrane vesicles, nanopods and/or nanotubes produced by hyperthermophilic archaea of the genus Thermococcus. Biochem Soc Trans. 2013;41(1):436–42.
Stetter KO. Hyperthermophilic microorganisms. Astrobiology: the quest for the conditions of life. 2002;169–84.
Kim KW. Electron microscopic observations of prokaryotic surface appendages. J Microbiol. 2017;55:919–26.
Moissl C, Rachel R, Briegel A, Engelhardt H, Huber R. The unique structure of archaeal ‘hami’, highly complex cell appendages with nano-grappling hooks. Mol Microbiol. 2005;56(2):361–70.
Liu J, Soler N, Gorlas A, Cvirkaite-Krupovic V, Krupovic M, Forterre P. Extracellular membrane vesicles and nanotubes in Archaea. MicroLife. 2021;2.
Hattori S. Syntrophic acetate-oxidizing microbes in methanogenic environments. Microbes Environ. 2008;23(2):118–27.
Morris BE, Henneberger R, Huber H, Moissl-Eichinger C. Microbial syntrophy: interaction for the common good. FEMS Microbiol Rev. 2013;37(3):384–406.
Schink B. Energetics of syntrophic cooperation in methanogenic degradation. Microbiol Mol Biol Rev. 1997;61(2):262–80.
van Wolferen M, Orell A, Albers S-V. Archaeal biofilm formation. Nat Rev Microbiol. 2018;16(11):699–713.
Fröls S, Dyall-Smith M, Pfeifer F. Biofilm formation by haloarchaea. Environ Microbiol. 2012;14(12):3159–74.
Abisado RG, Benomar S, Klaus JR, Dandekar AA, Chandler JR. Bacterial quorum sensing and microbial community interactions. MBio. 2018;9(3):e02331-e2417.
Falà AK, Álvarez-Ordóñez A, Filloux A, Gahan C, Cotter PD. Quorum sensing in human gut and food microbiomes: Significance and potential for therapeutic targeting. Front Microbiol. 2022;4389.
• Prescott RD, Decho AW. Flexibility and adaptability of quorum sensing in nature. Trends Microbiol. 2020;28(6):436–44. The authors highlight the flexibility of QS in allowing bacteria to respond to changing environmental conditions, including changes in nutrient availability and temperature. The paper also discusses the ability of QS to adapt and evolve in response to selective pressures, such as exposure to antibiotics or competition with other microorganisms. The authors highlight the potential for QS to be manipulated for therapeutic purposes, such as through the development of QS inhibitors.
Zhang G, Zhang F, Ding G, Li J, Guo X, Zhu J, et al. Acyl homoserine lactone-based quorum sensing in a methanogenic archaeon. ISME J. 2012;6(7):1336–44. https://doi.org/10.1038/ismej.2011.203.
Liao Y, Williams T, Ye J, Charlesworth J, Burns B, Poljak A, et al. Morphological and proteomic analysis of biofilms from the Antarctic archaeon. Halorubrum lacusprofundi Sci Rep. 2016;6(1):1–17.
Paggi RA, Martone CB, Fuqua C, De Castro RE. Detection of quorum sensing signals in the haloalkaliphilic archaeon Natronococcus occultus. FEMS Microbiol Lett. 2003;221(1):49–52.
Tommonaro G, Abbamondi GR, Iodice C, Tait K, De Rosa S. Diketopiperazines produced by the halophilic archaeon, Haloterrigena hispanica, activate AHL bioreporters. Microb Ecol. 2012;63:490–5.
• Zhang L, Li S, Liu X, Wang Z, Jiang M, Wang R, et al. Sensing of autoinducer-2 by functionally distinct receptors in prokaryotes. Nature Commun. 2020;11(1):5371. The authors used a combination of bioinformatics, genetic manipulation, and biochemical assays to identify and characterize three functionally distinct types of AI-2 receptors in bacteria. These receptors are found in different microbial species and have different structural and functional properties, allowing them to sense and respond to AI-2 in different ways. The paper demonstrates the potential for these receptors to be targeted for the development of new antibacterial therapies.
Nichols JD, Johnson MR, Chou C-J, Kelly RM. Temperature, not LuxS, mediates AI-2 formation in hydrothermal habitats. FEMS Microbiol Ecol. 2009;68(2):173–81.
Paggi RA, Madrid EA, D’Alessandro CP, Cerletti M, De Castro R. Growth phase-dependent biosynthesis of Nep, a halolysin-like protease secreted by the alkaliphilic haloarchaeon Natrialba magadii. Lett Appl Microbiol. 2010;51(1):36–41.
Grandclément C, Tannières M, Moréra S, Dessaux Y, Faure D. Quorum quenching: role in nature and applied developments. FEMS Microbiol Rev. 2016;40(1):86–116.
Sikdar R, Elias M. Quorum quenching enzymes and their effects on virulence, biofilm, and microbiomes: a review of recent advances. Expert Rev Anti Infect Ther. 2020;18(12):1221–33.
Ng FS, Wright DM, Seah SY. Characterization of a phosphotriesterase-like lactonase from Sulfolobus solfataricus and its immobilization for disruption of quorum sensing. Appl Environ Microbiol. 2011;77(4):1181–6.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no conflict of interest.
Human and Animal Rights and Informed Consent
N/A
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Weiland-Bräuer, N. Symbiotic Interactions of Archaea in Animal and Human Microbiomes. Curr Clin Micro Rpt 10, 161–173 (2023). https://doi.org/10.1007/s40588-023-00204-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40588-023-00204-7